Three Genealogy Conferences I Wish I Could Have Gone To - Mon, 26 May 2025
Between 2012 and 2018, I attended 11 international genealogy conferences.
They included:
- 3 times RootsTech in Salt Lake City (2012, 2014 and 2017)
- 2 Unlock the Past genealogy cruises around Australia and New Zealand (2013 and 2016)
- the Geanovium genealogy technology conference in Leiden, the Netherlands (2014)
- the Ontario Genealogy Society conference in Toronto (2016)
- the IAJGS (International Association of Jewish Genealogy Societies) conference in Disney World, Florida (2017)
- the Great Canadian Genealogical Summit in Halifax, Nova Scotia (2017)
- the Family Tree DNA Genetic Genealogy Conference in Houston (2017)
- the KDGS (Kelowna District Genealogical Society) conference in Kelowna, British Columbia in 2018.
I was a speaker at all these conferences. Each one was unique and wonderful in its own way. Meeting in person with genealogy friends who until then I had only known online was the highlight of each. And many of these friends attended several of these conferences.
There are 3 more conferences that I had made plans to go to and was looking forward to. I wish I could have gone to these as well.
1. The Manitoba Genealogy Society’s 1st Provincial Conference.
The MGS is my home province’s genealogy society. I have spoken at a few local meetings for them before, but this was going to have been their first attempt at a bigger event of maybe 400 attendees. It was going to have taken place from June 12 to 14, 2020 in Winnipeg where I live.
I was scheduled to speak at the event and I know that Diahan Southard was being brought in as the keynote speaker. I was looking forward to meeting Diahan and spending time with her, and was hoping that I could arrange to pick her up at the airport and wine and dine her a bit.
Unfortunately, on November 8, 2019, the MGS announced that they had to cancel the Conference. As it turns out, the Conference likely would have been cancelled the next year anyway due to Covid.
2. The MyHeritage LIVE 2020 conference.
In December 2019, MyHeritage announced its 3rd MyHeritage LIVE conference. The previous two were held in Oslo, Norway in 2018 and in Amsterdam, the Netherlands in 2019.
But this one would be special because it was to be held in Tel Aviv, Israel. Since that’s where the MyHeritage headquarters is, what it means is that many more of their staff would be able to participate, and attendees would be able to visit the company offices.
I submitted my application to be a speaker. I had never been to Israel myself and my wife and I were looking forward to touring the country for two weeks before or after the conference.
Alas this conference was a casualty of Covid.
I’m hoping now that the major Covid worries seem to have passed, that MyHeritage will resurrect their own MyHeritage LIVE conferences and maybe one day when the situation in Israel is better resolved, they will announce a conference in Israel, and I will be sure to plan to attend.
3. The Find Your Roots River Cruise
This one was planned in 2022, scheduled for 2024 with the hope that Covid worries would by then be no more. It was to have been a genealogy European river cruise with headliners Blaine Bettinger and Judy Russell.
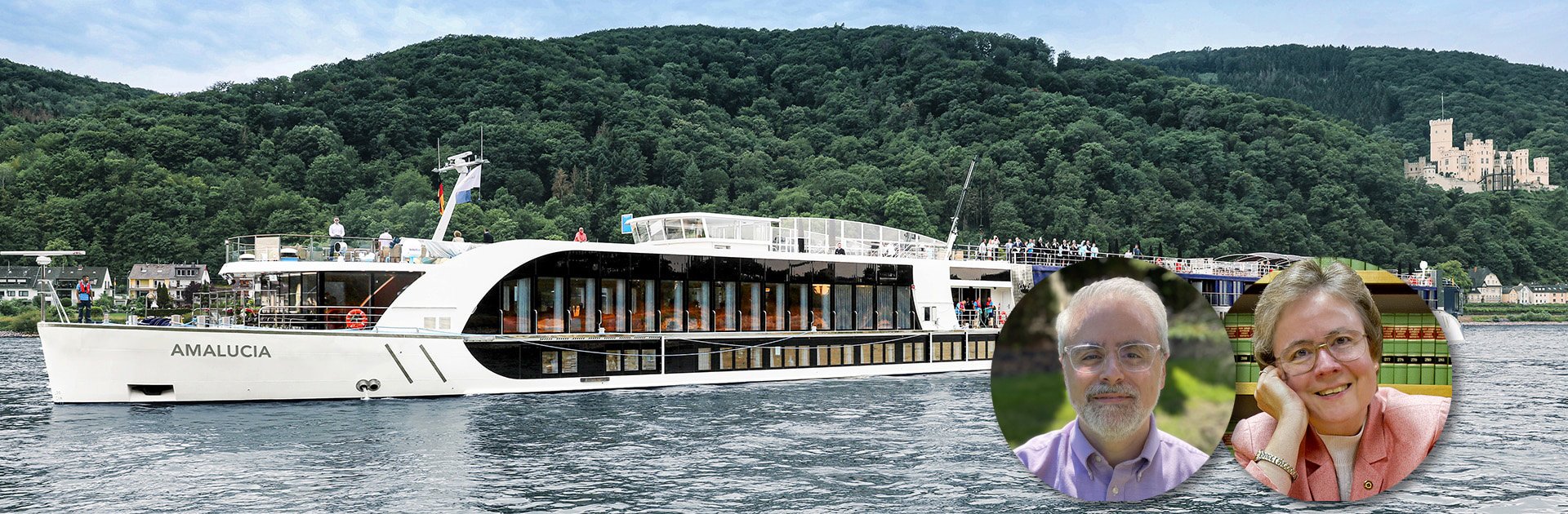

They were to be the only speakers, so for once I’d be able to just enjoy Blaine and Judy’s talks and not have any obligations. My wife and I were very much looking forward to this cruise and spending time with a shipfull of genealogy enthusiasts.
I wrote about this cruise in a 2022 blog post: Genealogy River Cruising in 2024
Unfortunately, this cruise was cancelled in 2023 as they did not attract enough people to fill the ship, with the price of the cruise being fairly high.
The Future
I’m not sure how many more conferences I will attend in person. Since Covid, so much happens virtually now. But if there is something new and special, or a person I need to meet, then I’ll certainly consider going.
What genealogy conferences do you wish you could have gone to?







 Feedspot 100 Best Genealogy Blogs
Feedspot 100 Best Genealogy Blogs





