MyHeritage has a very nice feature in their chromosome browser. If three or more people all triangulate over a segment, the browser will place a rectangle with rounded corners over the triangulating segment for you. These form a triangulation group. All of the people you select must all match each other on that segment for the rectangle to show. If just one person doesn’t match all the others, then the rectangle will not show.

I have been having a difficult time with my own matches at MyHeritage. The reason is that out of my 12,142 DNA Matches that I have there, I only have my uncle and one other person that I know how I’m related or who is our MRCA (Most Recent Common Ancestor).
At MyHeritage, my Uncle matches me on 52 segments that total 1994.1 cM. My next highest match is someone sharing 9 segments with me totaling 141.4 cM. MyHeritage estimates him as a 1c2r to 2c1r, but I know that he is more distant than that or I would have already been able to place him in my tree.
I uploaded my uncle’s Family Tree DNA test to MyHeritage. But because our common ancestor is my father’s parents, all his DNA will allow me to do is to help separate segment matches on my father’s side from my mother’s. Segment matches triangulating with my uncle and myself almost always will be my father’s side. Missing B-C segment matches over those triangulating segments (i.e. I match my uncle and Person C, but my uncle does not match Person C) will almost always be on my mother’s side.
My Other Known Relative
I have one other person at MyHeritage that I match to whose relationship I know. It is a 2c2r who is my father’s mother’s father’s mother’s brother’s son’s son and his MRCA would be FMFMR in the notation DMT uses. We share 8 segments totaling 79.4 cM.
When I use the MyHeritage chromosome browser with my uncle and this cousin, I have a problem:

You can see how nicely MyHeritage has boxed 4 triangulations between myself and my uncle (red) and my cousin (yellow) on chromosomes 1, 2, 8 and 18.
There is one small segment on chromosome 4 where I match my cousin but not my uncle. That is fine. I could have my uncle’s mother’s father segment there, but my uncle could have his mother’s mother’s segment.
The problem occurs on chromosomes 4, 11 and 16, segments where I match both my uncle and my cousin, but MyHeritage does not show a triangulation box. That includes a large match of 15.4 cM with my cousin on chromosome 4. These are Missing B-C matches. I cannot match both my uncle and my cousin without my uncle also matching my cousin. Therefore on these three segments, either I must be matching my third cousin on my mother’s side through some unknown relationship, or these are by-chance false matches between me and my cousin where either of his chromosomes is matching either of mine over the segment.
That throws a wrench in the works as far as assigning an MRCA to my cousin in Double Match Triangulator. If I do, DMT will assume the ancestral path FMFMR would apply to all 8 segments. If I had a lot of other people with known MRCAs, this wouldn’t be too much of a problem, because DMT’s “majority rule” would mitigate the few incorrect assignments using the majority of correct assignments. But since I don’t have any other people to use, I know DMT will assign my FMFMR side to the other three segments, and those incorrect assignments will replicate when DMT iterates its clustering of matches and recompution of the ancestral paths.
Confirming the Triangulation Group at MyHeritage
My cousin does not match my uncle on 3 segments. Maybe if I can identify the people in the triangulation groups of those three segments, I can have more information to investigate to determine if those segments are indeed on my mother’s side.
So I’ll first look at my triangulations on that large 15.4 cM segment on chromosome 4. But how do I do that at MyHeritage? MyHeritage allows you to select up to 7 people to compare. However they give you no way to know what people you need to pick to match on that specific segment.
This is where Double Match Triangulator comes to the rescue. I don’t have my cousin’s segment match file from MyHeritage. But I do have my own and my uncle’s. Those two are enough.
I start DMT, setting my segment match file as File A and my uncle’s as File B. I open my People file and set my uncle’s MRCA to FR, and I run DMT.
DMT shows me all the people who triangulate with myself and my uncle and assign them all an ancestral path and cluster of F (Father). These are true triangulations, since my file tells me that I match Person C and my uncle, and my uncle’s file tells me that he matches Person C.
DMT also shows me the people who have Missing B-C matches (that my uncle does not match to on that segment) and put them in the cluster of M (mother).
Now I look at the DMT output and go down to chromosome 4 in M (mother) section and look at the Missing B-C matches between 58 Mbp and 77 Mbp. I’m adding a box around this range to visually denote them.
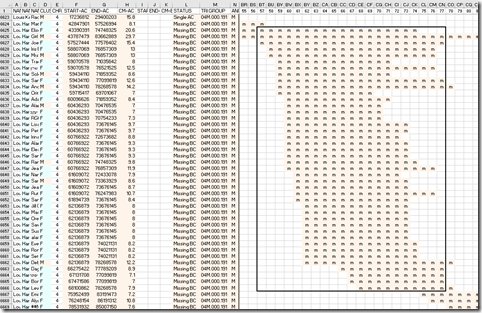
There are 39 people that I match to on this segment that are Missing B-C matches. Theoretically, all these people should form a triangulation group here, since they all triangulate over the same region. I know they all match both me and my uncle, but what I don’t know is if they match each other. I would expect they would since I have found that triangulations or Missing B-C matches of 7 cM or more at other companies generally are all valid and do match each other.
So now I’ll use MyHeritage’s nice triangulation display to verify that these matches are in this maternal triangulation group.
I’ll start by adding the my cousin, and the first Missing B-C person

That’s good. MyHeritage gives the rectangle so it triangulates.
Now I’ll add the the next person. I know exactly who to add because DMT gives me each person’s exact names. Then I can simply copy from the DMT spreadsheet and paste into MyHeritage’s Chromosome Browser search field and presto, that person is found. This is especially useful when the name is in a foreign script like Cyrillic or Hebrew.
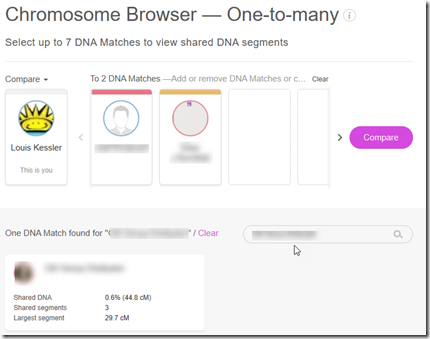
I add that found match and click compare and I get:
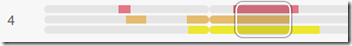
I add the next two and I still have a triangulation group:
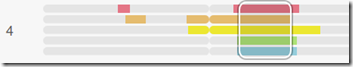
This happens with the next person I add::
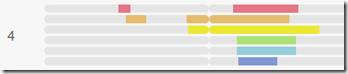
MyHeritage no longer draws the triangulation box. This last person in dark blue only has an 8 cM match with me, and MyHeritage has proven the match to not match everyone in the group.
If I had added added 7 people at once and there was no triangulation box, I wouldn’t know who is breaking the triangulation group. If that happens, it is best to add people one at a time, and verify the triangulation group for each one. If the triangulation group breaks, you know who it is who is out.
So now I note that the above person is not in the group, delete them and try the next person.
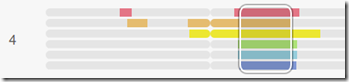
Aha! That’s better! Once the 7 slots on MyHeritage fills up, I just can delete the last person and replace with the next and thus test all 37 people reasonably quickly. It does take a few seconds for MyHeritage to compare 8 people because there are 8 x 7 / 2 = 28 comparisons for them to make and they are comparing all segments on all chromosomes each time. I’m so glad MyHeritage is correctly checking all 28 comparisons rather than incorrectly just doing the 8 comparisons of 1 vs 2, 2 vs 3, … 7 vs 8. It is possible that 3 might match 4 but not match anyone else.
As I went along, every other person was turning up false. I was quite shocked to end up with only the above 6 of the 41 people being in the triangulation group. These are all segments above 7 cM. Those rejected went as high as 12.2 cM.
This prompted me to do the same for the largest triangulating match with my cousin on my father’s side. This is by coincidence also a 15.4 cM match on chromosome 2 between 221 Mbp and 234 Mbp. Here’s the section of the DMT report for the triangulating segments with a box around the 56 matches in that range.
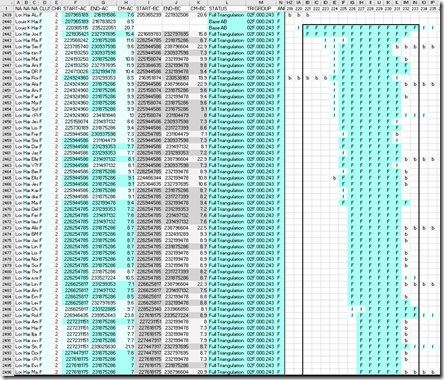
An “F” in darker blue is the part of the segment that triangulates on my father’s side. An “f” in lighter blue is where I match the person but my uncle does not. A “b’ in grey is where my uncle matches the person but I do not.
I checked many of them one-by-one using MyHeritage’s chromosome browser, and only found 3 out of the 56 matches to form a triangulation group with my uncle and myself.

That included my cousin 15.4 cM and the two large matches of 23.8 cM and 21.9 cM that you can see near the bottom of the DMT report which are extending to the right with ‘f’s in lighter blue. All the other 53 must be false matches.
Explaining False Triangulating Segments
In all the work that I have done up to now, I have come to agree with the conclusion of Jim Bartlett that almost all segment matches 7 cM or more that triangulate are valid and will form a triangulation group. I have extended that thinking to include Missing B-C matches 7 cM or more, as long as B-C do share segments elsewhere which tells you that the B-C segment is an explicit non-match
But here at MyHeritage, I have many triangulating and Missing B-C matches above 7 cM that don’t match with the triangulation group. One was even as large as 16 cM. The sample of them that I checked on MyHeritage gave me this:
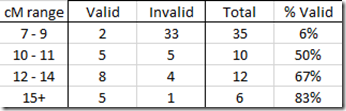
This is not a good result for MyHeritage, and indicates to me that their segment matching is less reliable than other companies and can produce many false triangulations above 7 cM.
If I could hazard a guess to why, it might be because of the imputation and stitching techniques that they use to determine matching segments. That technique may be less precise than what the other companies use. Others, including Roberta Estes have less faith in imputation techniques.
Conclusion
The display of triangulation groups in MyHeritage’s chromosome browser is a wonderful feature. It will help you ensure that overlapping segments all match each other forming a triangulation group that likely was passed down from a common ancestor.
But it also enabled me to illustrate that MyHeritage’s matching is not as reliable as other companies, since their match data allows the formation of invalid triangulations with larger segments than at other companies.
I recommend that Double Match Triangulator users, when using MyHeritage data, that you increase your “Min Triang” setting from 7 cM to at least 12 cM or 15 cM to ensure fewer false positives are included in your MyHeritage triangulations and Missing B-C matches.
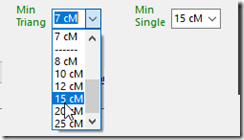
At some point in a future blog post, I’d do a full run of DMT with my MyHeritage segment match data.












 Feedspot 100 Best Genealogy Blogs
Feedspot 100 Best Genealogy Blogs





